
| Version | Change log |
| Cytoscape for Mac OS X 3.10.0 May 12, 2023 | |
| Cytoscape for Mac OS X 3.9.0 Oct 12, 2021 |

Total downloads
47
Last month's downloads
0
Last week's downloads
0
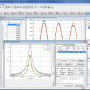
... the highly rated data analysis software "MagicPlot Pro for Mac OS X" developed by MagicPlot.com. This impressive software is a must-have tool for researchers, scientists, engineers, and students who want to transform their data into detailed visualizations with ease and precision. The user-friendly interface and comprehensive ...
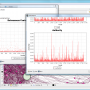
... one of the most innovative and powerful data visualization software packages available on Mac OS X – Gephi. Developed by Mathieu Bastian, this software ... of complex data sets, making it a must-have for businesses, researchers, and anyone who needs to make ...
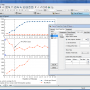
... a software developed by MagicPlot.com called MagicPlot Student for Mac OS X. This powerful tool is perfect for students and researchers who need to visualize and ... and user-friendly interface, MagicPlot Student makes it easy for users to create custom graphs, charts, and plots ...